GeT présent à JOBIM 2017
La plateforme GeT présente, à JOBIM 2017 à Lille, un poster sur les dernières analyses de comparaison de données Long Reads pour l’assemblage de novo de différents génomes (bactérie, champignon, tomate et poisson).
Résumé :
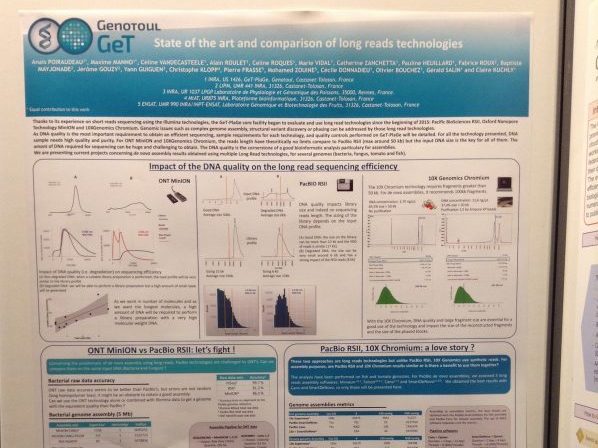
Thanks to its experience on short reads sequencing using the Illumina technologies, the GeT-PlaGe core facility began to evaluate and use long read technologies since the beginning of 2015: Pacific BioSciences RSII, Oxford Nanopore Technology MinION and 10XGenomics Chromium. Genomic issues such as complex genome assembly, structural variant discovery or phasing can be addressed by those long read technologies.
As DNA quality is the most important requirement to obtain an efficient sequencing, sample requirements for each technology, and quality controls performed on GeT-PlaGe will be detailed. For all the technology presented, DNA sample needs high quality and purity. For ONT MinION and 10XGenomics Chromium, the reads length have theoritically no limits compare to PacBio RSII (max around 50 kb) but the input DNA size is the key for all of them. The amont of DNA required for sequencing can be huge and challenging to obtain. The DNA quality is the cornerstone of a good bioinformatic analysis particulary for assemblies.
We are presenting current projects concerning de novo assembly results obtained using multiple Long Read technologies, for several genomes (bacteria, fungus, tomato and fish).
Télécharger le poster de GeT : Poster Get JOBIM2017